Objective
Determine
HIV drug resistance mutations (DRMs) prevalence at low and high levels in
ART-experienced patients experiencing virologic failure (VF).
Methods
29
subjects from 18 counties in Hunan Province that experienced VF were evaluated
for the prevalence of DRMs (Stanford DRMs with an algorithm value ≥15, include low-,
intermediate and high-level resistance) by both Sanger sequencing (SS) and deep
sequencing (DS) to 1% frequency levels.
Results
DS
was performed on samples from 29 ART-experienced subjects; the median viral
load 4.95×104 c/ml; 82.76% subtype CRF01_AE. 58 DRMs were
detected by DS. 18 DRMs were detected by SS. Of the 58 mutations detected by
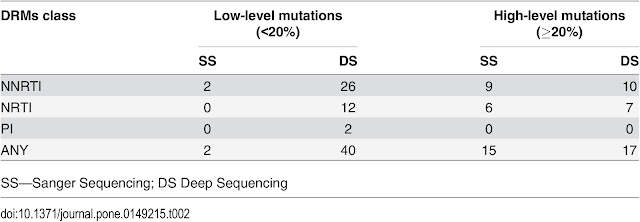
DS, 40 were at levels <20% frequency (26 NNRTI, 12 NRTI and 2 PI) and the
majority of these 95.00% (38/40) were not detected by standard genotyping. Of
these 40 low-level DRMs, 16 (40%) were detected at frequency levels of 1–4% and
24 (60%) at levels of 5–19%. SS detected 15 of 17 (88.24%) DRMs at levels ≥ 20%
that were detected by DS. The only variable associated with the detection of
DRMs by DS was ART adherence (missed doses in the prior 7 days); all patients
that reported missing a dose in the last 7 days had DRMs detected by DS.
Conclusions
DS of VF samples from treatment experienced subjects
infected with primarily AE subtype frequently identified Stanford HIVdb NRTI
and NNRTI resistance mutations with an algorithm value 15. Low frequency level
resistant variants detected by DS were frequently missed by standard genotyping
in VF specimens from antiretroviral-experienced subjects.
Below: Prevalence of ≥1% DRMs by Stanford HDRM (algorithm value ≥15)
Full article at: http://goo.gl/ZY39t9
By:
Xi Chen, Xiaobai Zou, Jianmei He, Jun Zheng
Hunan Provincial Center for
Disease Control and Prevention, Changsha, China
Jennifer Chiarella, Michael J. Kozal
Yale School of Medicine, New
Haven, United States of America
More at: https://twitter.com/hiv insight

No comments:
Post a Comment